 The locations of the fragments of DNA show up on the film as bands. As a result, shorter fragments migrate farther from the origin as they move through the gel.Īfter the gel is run, the DNA is labeled using a radioactive probe and the gel is exposed to x-ray film, which changes color in the presence of radioactivity. This distance is roughly proportional to the inverse of the fragment's length. In gel electrophoresis an electrical current is transmitted through the gel causing the fragments of DNA to migrate through the gel according to their electrophoretic mobility. After exposure to the same restriction enzymes, the resulting strands of DNA would be GC, GCAAGGCGAA and TTCGCCC.Īfter exposure to the restriction enzymes, the two mixtures are transferred to a gel and electrophoresis is performed. This individual has a point mutation so that their DNA sequence is GCGCAAGGCGAATTCGCCC. Next, consider a sample of the same region of DNA from a second individual. The resulting strands from this RFLP would be GC, GCAAGGCGAA, TTCGC, and CG. EcoRI recognizes the sequence GAATTC and it cleaves the bond between the adenine and the thymine. As discussed, HaeIII cleaves between C and G on the sequence GCGC. The restriction enzymes HaeIII and EcoRI are both added to the mixture. For example, consider the strand of DNA from one individual with the sequence GCGCAAGGCGAATTCGCGC. This amplified DNA is then combined with a set of restriction enzymes, which cleave the DNA in specific locations. When performing RFLP, the target DNA is usually subjected to polymerase chain reaction, which produces millions of copies of strands of DNA identical to the original. Each of these enzymes cleaves DNA between different, and specific, sequences of nucleotides. Over 90 different restriction enzymes have been isolated from different species of bacteria. Bacteria naturally produce restriction enzymes and they use them to cleave the DNA from foreign organisms. For example, the restriction enzyme HaeIII recognizes the sequence GCGC and it cleaves the bond between middle cytosine and guanine. This is simply the replacement of a single nucleotide by a different one.Ī special type of protein called a restriction enzyme, or a restriction endonuclease, can recognize specific sequences of nucleotides on DNA and then cleave the DNA at these locations. Two families of these repeats are found quite often in DNA: variable number of tandem repeats (VNTRs) and short tandem repeats (STR). In vertebrates (animals with a backbone), there are regions of DNA that contain many repetitions of the same sequence. Deletions are regions where nucleotides have been removed. Insertions are regions of DNA where nucleotides have been added to a sequence. There are a variety of types of mutations in DNA. Differences in individuals result from small variations, called mutations, in the sequence of DNA. The sequence of these nucleotides is extremely important, as it determines the structure of all of the molecules in an individual. 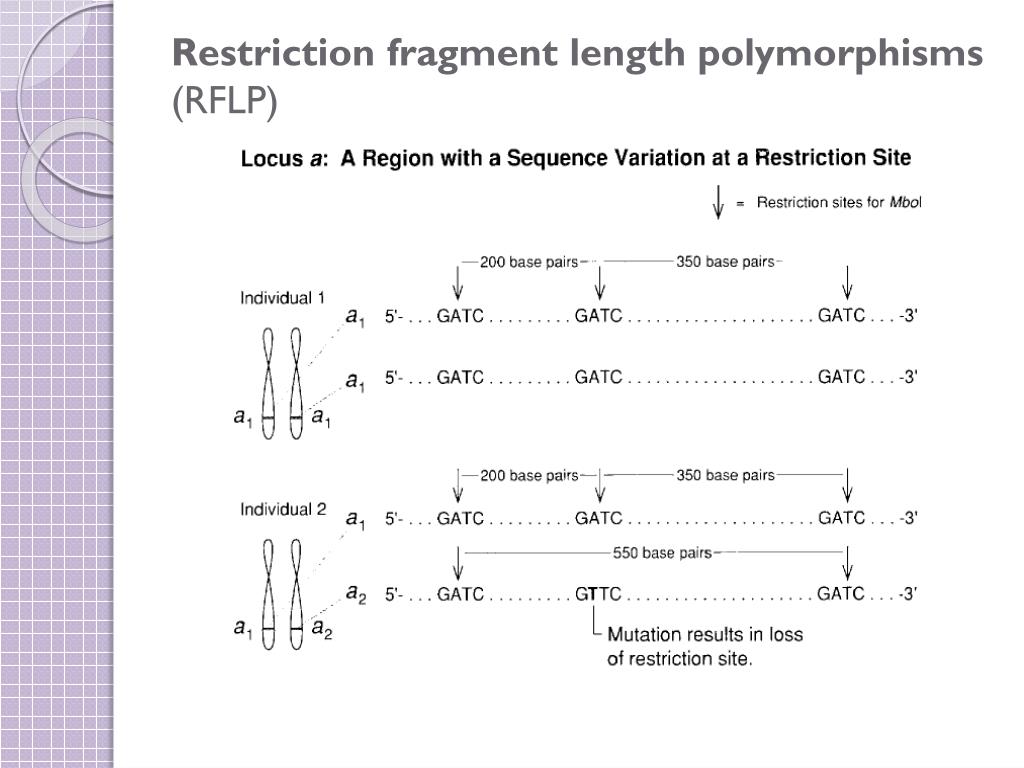
The four nucleotides are adenine (A), guanine (G), cytosine (C), and thymine (T). The DNA molecule is made up of a sequence of four smaller molecules called nucleotides. RFLP is used as part of DNA fingerprinting, to detect genetic diseases and to determine genetic relationships between species. 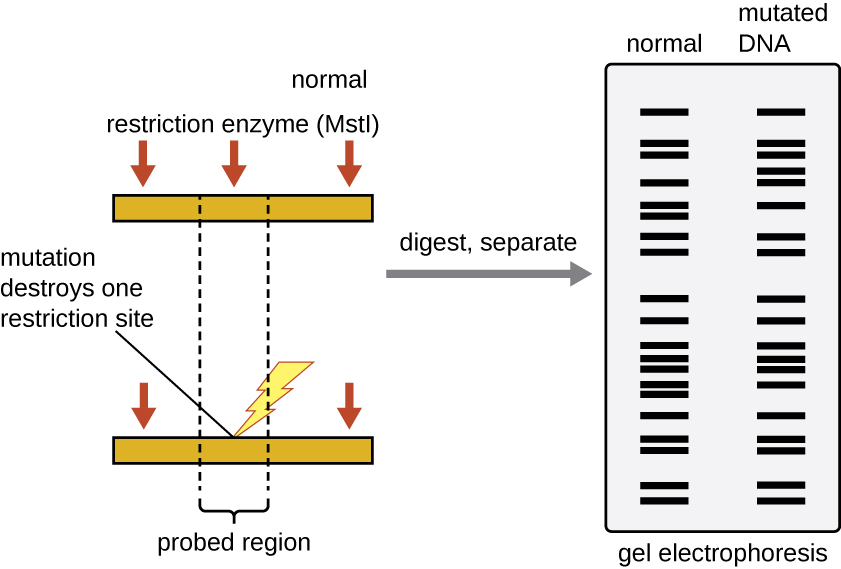
Electrophoresis is then used to separate the strands according to their length. Mutations within the DNA result in strands of different lengths. Special enzymes that cleave the DNA in specific locations are used to digest strands of DNA. RFLP, or restriction fragment length polymorphism, is a molecular biological technique used to compare DNA from two samples. RFLP (Restriction Fragment Length Polymorphism) 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
